Federated one-shot likelihood pooling
poolLikelihoodProfiles.RdCombines profile log-likelihood vectors from multiple federated sites by pointwise summation. This is the core of the one-shot federated analysis: summing log-likelihoods is equivalent to multiplying likelihoods, yielding the joint likelihood under independence across sites.
Before summing, the function validates that all sites used identical beta grid vectors. If grids differ, linear interpolation to a common grid over the overlapping beta range is applied with a warning. Pooling never extrapolates beyond a site's evaluated range.
Arguments
- siteProfileList
Named list of site profile objects, each the output of
fitOutcomeModel.- validateGridAlignment
Logical. If TRUE (default), checks that all sites used the same betaGrid and warns if interpolation is needed.
Value
A list with class "medusaCombinedProfile" containing:
- betaGrid
The common beta grid used for pooling.
- logLikProfile
Numeric vector of summed log-likelihood values.
- siteContributions
Data frame with siteId, nCases, nControls, and diagnostic flags per site.
- nSites
Number of sites pooled.
- totalCases
Total number of cases across sites.
- totalControls
Total number of controls across sites.
Details
Pool Profile Log-Likelihoods Across Sites
The pooling operation is mathematically straightforward: for each grid point \(b_k\), the combined log-likelihood is $$\ell_{combined}(b_k) = \sum_{s=1}^{S} \ell_s(b_k)$$ where \(\ell_s\) is the profile log-likelihood from site \(s\).
This pooling is exact under the assumption that observations are independent across sites and every site uses the same beta grid. If interpolation is needed, pooling remains a close numerical approximation on the common grid, but only over the beta range evaluated by every site. No iterative communication protocol is needed: each site computes its profile once and shares only the numeric vector.
References
Luo, Y., et al. (2022). dPQL: a lossless distributed algorithm for generalized linear mixed model with application to privacy-preserving hospital profiling. Journal of the American Medical Informatics Association, 29(8), 1366-1373.
Examples
profiles <- simulateSiteProfiles(nSites = 3, trueBeta = 0.5)
combined <- poolLikelihoodProfiles(profiles)
#> Pooling profile likelihoods from 3 site(s)...
#> Pooling complete: 3 sites, 402 total cases, 5598 total controls.
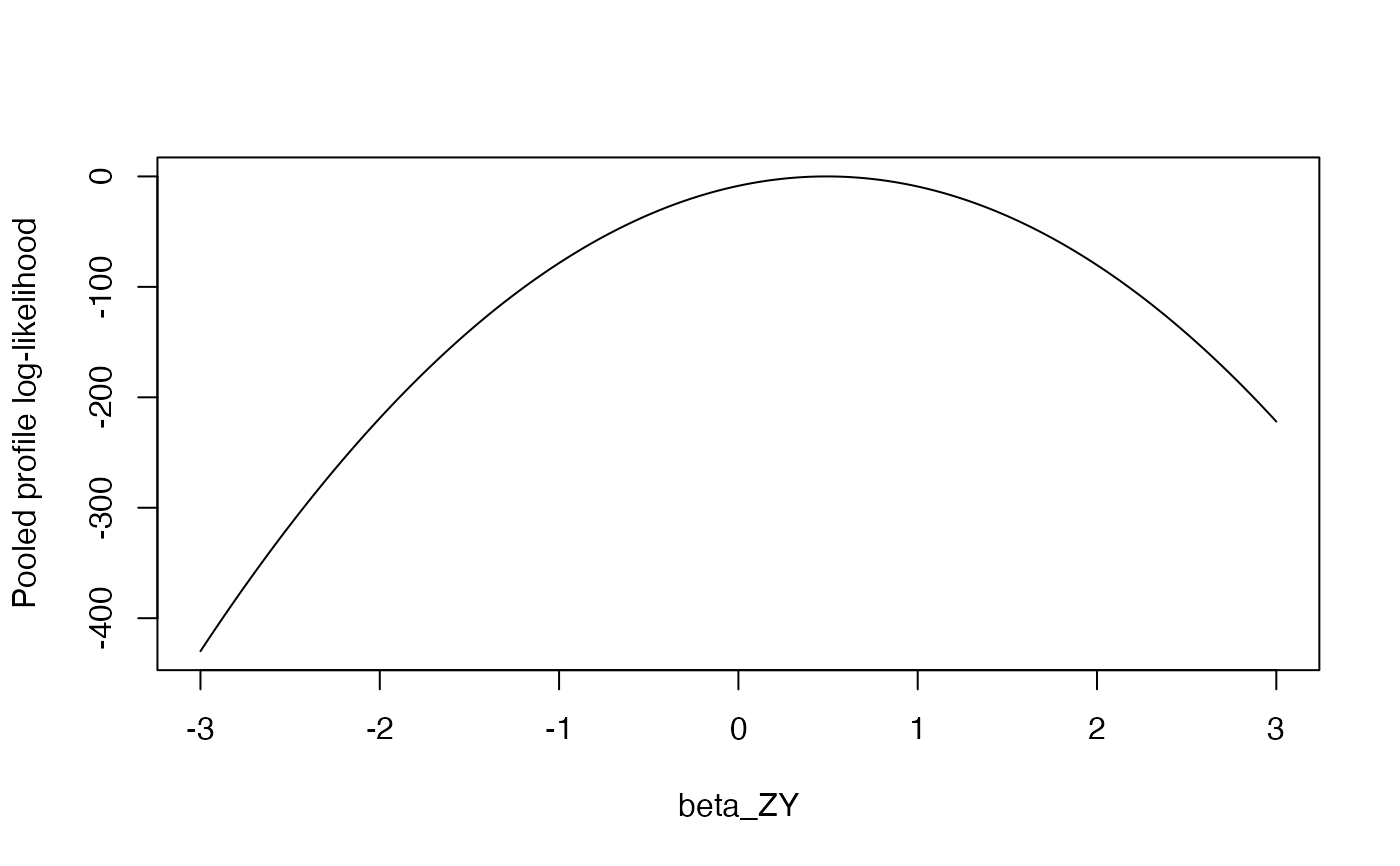
plot(combined$betaGrid, combined$logLikProfile, type = "l",
xlab = "beta_ZY", ylab = "Pooled profile log-likelihood")