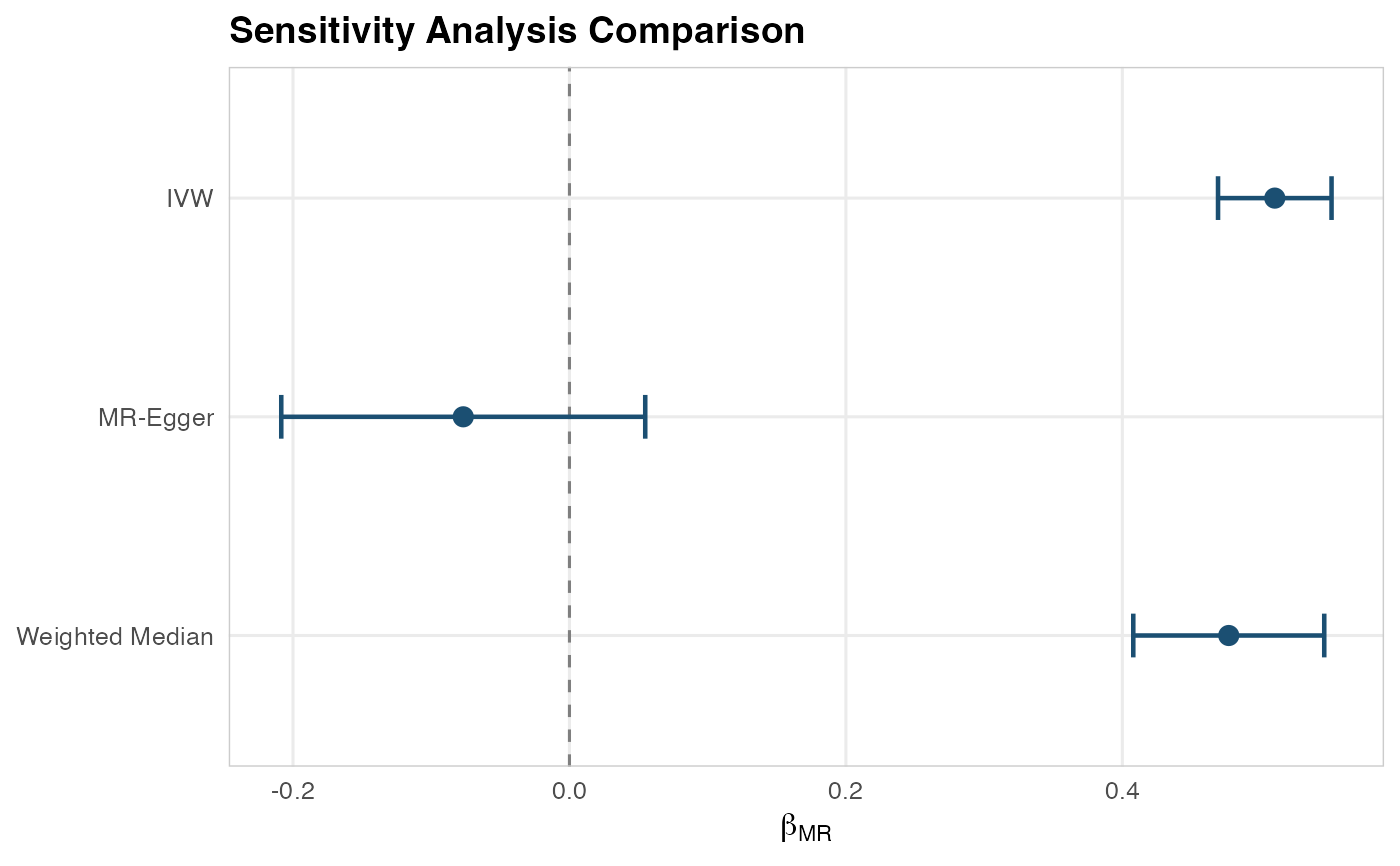
Forest plot comparing MR methods
plotSensitivityForest.RdCreates a horizontal forest plot showing estimates from all sensitivity analysis methods side by side with confidence intervals.
Arguments
- sensitivityResults
Output of
runSensitivityAnalyses.
Examples
set.seed(42)
nSnps <- 10
perSnp <- data.frame(
snp_id = paste0("rs", 1:nSnps),
effect_allele = rep(c("A", "C", "G", "T", "A"), length.out = nSnps),
other_allele = rep(c("C", "G", "T", "A", "C"), length.out = nSnps),
eaf = seq(0.1, 0.7, length.out = nSnps),
beta_ZY = rnorm(nSnps, 0.15, 0.02),
se_ZY = rep(0.02, nSnps),
beta_ZX = rnorm(nSnps, 0.3, 0.05),
se_ZX = rep(0.05, nSnps)
)
results <- runSensitivityAnalyses(perSnp, engine = "internal")
#> Running sensitivity analyses with 10 SNPs...
#> Engine: internal
#> IVW...
#> MR-Egger...
#> Weighted Median...
#> Steiger filtering...
#> Warning: Steiger filtering is not implemented for binary outcomes because logistic-regression coefficients cannot be converted to correlations without additional scale assumptions. Returning NA result.
#> Leave-One-Out...
#> Sensitivity analyses complete.
plotSensitivityForest(results)
#> `height` was translated to `width`.